Cancer subtyping using cNMF and literature markers
An example of usage of cNMF to identify cancer subtypes
Purpose of this tutorial
Cancer is a complex disease that arises from the accumulation of genetic mutations. These mutations are vast, and differ from one patient to another. This makes it so that cancer is generally seen as a heterogeneous disease. This heterogeneity is reflected in the different subtypes of cancer, which are defined by the mutations that are present in the tumor. These subtypes generally have different clinical features, such as the presence of different mutations, the size of the tumor, and the presence of certain genes (key transcription factors). Additionally, these subtypes can be associated with different responses to treatment. Thus, understanding the molecular biology of different cancer types is key to developing effective treatments and, ultimately, improving patient outcomes. Here, I want to show a brief example of how we can use methods such as cNMF to identify cancer subtypes, particularly in Small-cell lung cancer (SCLC).
What do we know about SCLC?
While this isn’t intended to be a comprehensive overview of SCLC, it is important to understand what we already know about the disease, in order to subtype it.
SCLC has long been recognized as one of the most aggressive and heterogeneous cancers. For decades, researchers tried to make sense of this complexity by dividing tumors into “classic” and “variant” forms. But as molecular profiling advanced, a more refined classification emerged: one that highlights the role of dominant transcription factors in shaping tumor identity and behavior.
Thus, in this tutorial, we can define molecular subtypes as being biologically consistent groups of tumors characterized by three key features:
- a dominant transcriptional regulator or signaling pathway,
- a relatively stable molecular signature across different disease stages and experimental models,
- clear association with distinct biological behaviors or therapeutic responses observed in clinical settings
Today, four major molecular subtypes are recognized:
- SCLC-A: Driven by ASCL1, accounting for about half of all cases.
- SCLC-N: Defined by NEUROD1, often linked to variant morphology and poorer prognosis.
- SCLC-P: Characterized by POU2F3, representing non-neuroendocrine tumors with tuft cell features.
- SCLC-I: The newest player, marked by an inflamed phenotype with immune-related activity.
What about SCLC-Y?
Before SCLC-I was proposed, some researchers suggested a fourth subtype called SCLC-Y, supposedly driven by YAP1. YAP1 is a transcriptional co-activator involved in proliferation, survival, and drug resistance. Early studies hinted that it might define a distinct group of tumors. However, subsequent work revealed that YAP1 expression was inconsistent and often low, failing to meet the criteria for a stable, subtype-defining regulator. In other words, YAP1 activity was biologically relevant, but it didn’t reliably carve out a separate molecular identity for SCLC tumors.
The inflamed alternative: SCLC-I
In 2021, Gay et al.
For this tutorial, we will attempt to replicate the subtyping done in Gay et al. using a dataset from George et al. (2015) which can be found here. This same dataset was used by Gay et al to perform their subtyping.
Here is a summary of what we know about SCLC, from a slide I have created and use to present at conferences:
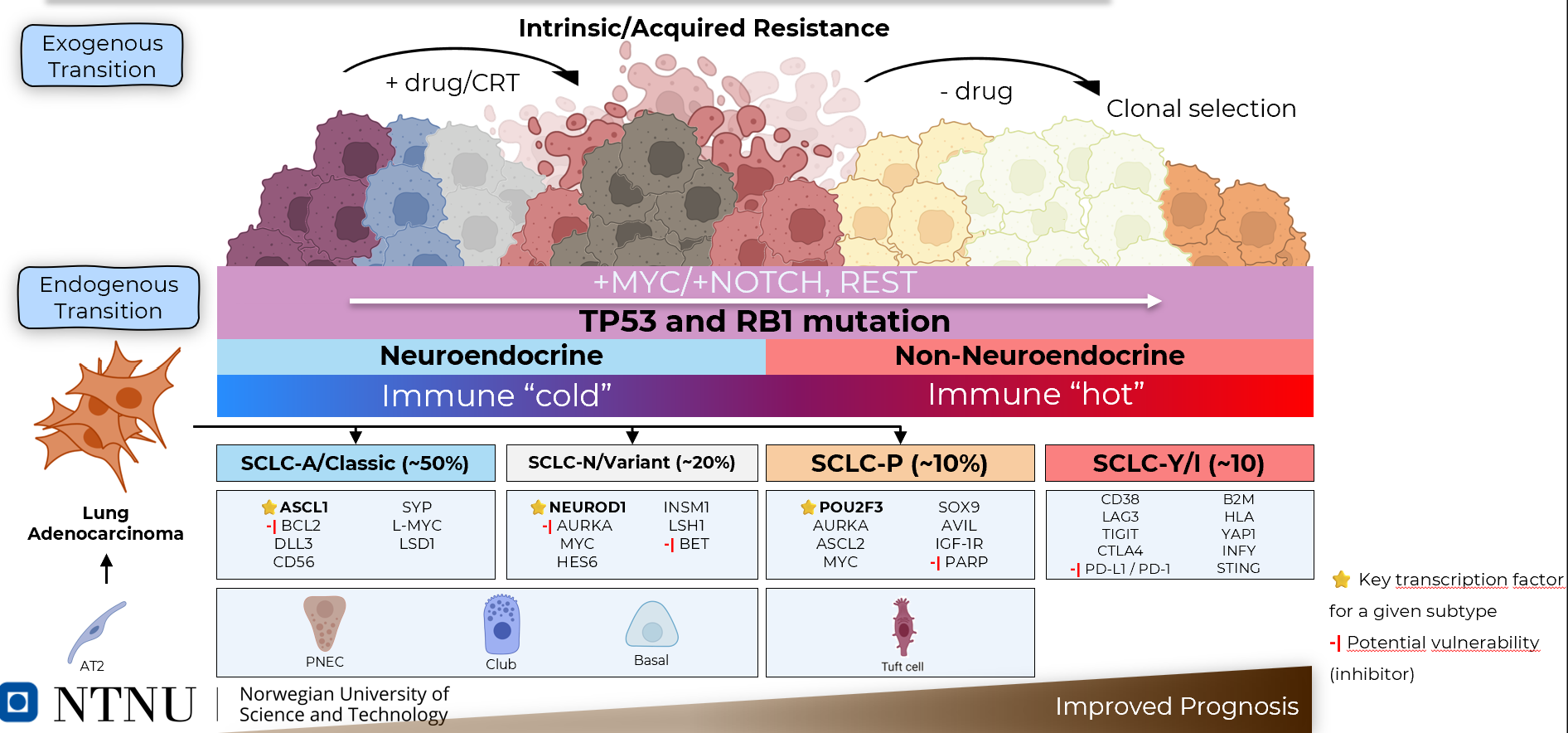
What is cNMF?
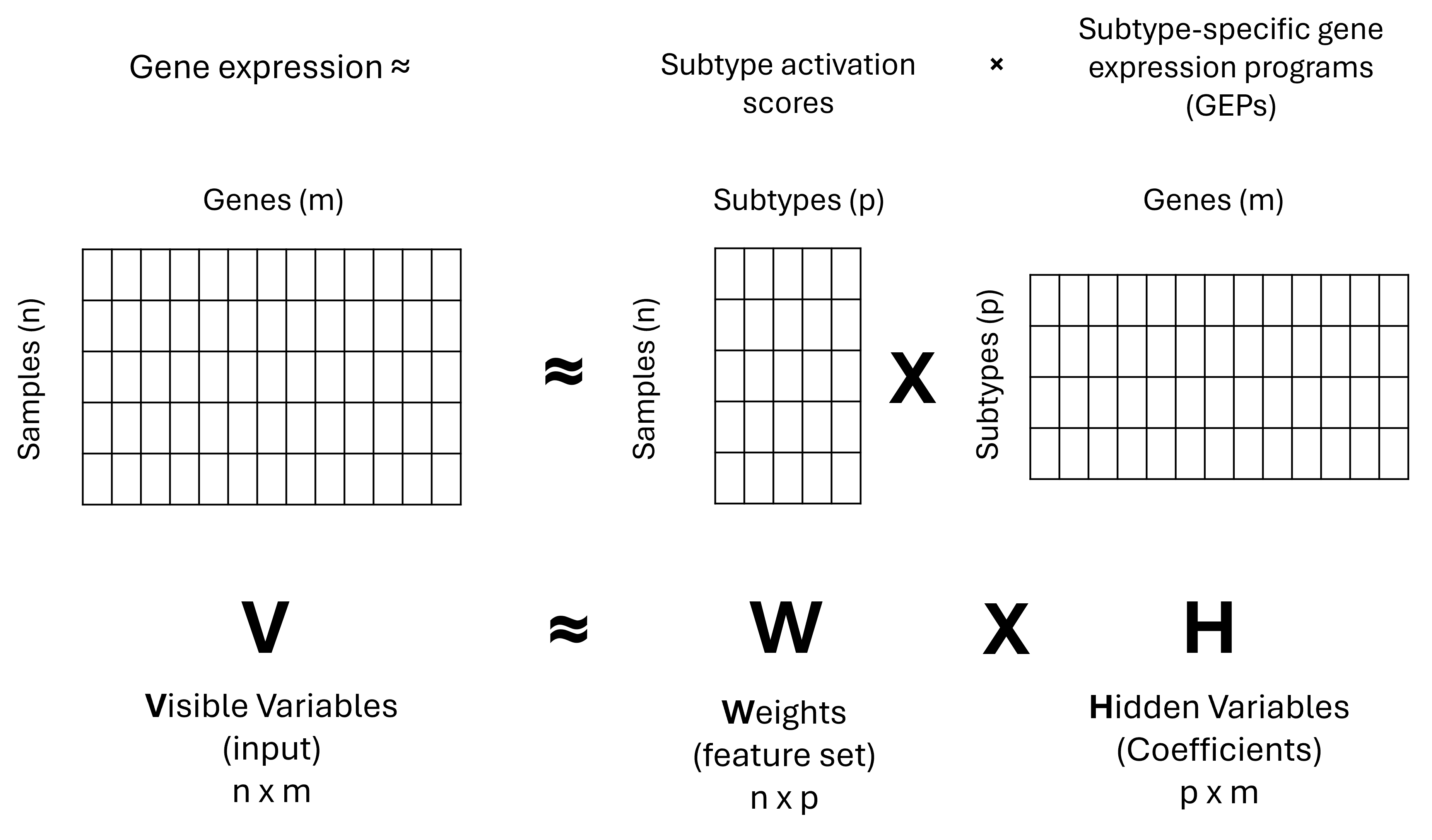
cNMF is a non-negative matrix factorization (NMF) method that is particularly useful for identifying cancer subtypes. The “c” in cNMF stands for “consensus” which means that the algorithm will run the analysis multiple times and find the most stable, reliable patterns.
cNMF can be used in a variety of settings, including single-cell RNA (scRNA) in order to look for groups of genes that work together, called gene expression programs. This can be extended to bulk RNA, this time looking for cancer subtypes (here we are talking about samples and not single cells).
The strength of cNMF is in its ability to solve compositional problems. Imagine sitting in a concert hall, listening to an orchestra. If you focus carefully, you start to notice recurring harmonies from the different instruments. That’s exactly what cNMF does with gene expression data. It picks a part the recurring harmonies of genes, the programs that consistently play together. These GEPs (gene expression programs) reveal the functional states of cells or the molecular identity of tumors, turning noise into meaningful biological insight.
In this tutorial, we will be using the python cNMF package which although has been used for scRNA, can also be used for bulk RNA data.
Reading the data
First, let’s import the necessary libraries.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
from cnmf import cNMF
from scipy.stats import zscore
from matplotlib.colors import rgb2hex
import scanpy as sc
import seaborn as sns
import os
from pathlib import Path
import pandas as pd
import anndata as ad
import numpy as np
plt.style.use("bmh")
plt.rcParams.update({"figure.figsize" : (12, 8),
"axes.facecolor" : "white",
"axes.edgecolor": "black"})
Now, lets read the count data. In this case, I am already providing the TPM counts, which were converted from FPKM (original data can be found here ). Check this repository link to find all the data necessary to follow this tutorial .
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
try:
_base_dir = Path(__file__).resolve().parent
except NameError:
_base_dir = Path.cwd()
data_file = _base_dir / "data" / "files" / "tpm_counts.csv"
count_data = pd.read_csv(data_file,sep="\t",
index_col=0,)
clinical_metadata_path = _base_dir / \
/ "data" / "files" / "data_clinical_patient.txt"
clinical_metadata = pd.read_csv(clinical_metadata_path, sep="\t")
clinical_metadata = clinical_metadata.iloc[4:, :]
# Subset clinical metadata with the samples from count_data
clinical_metadata_subset = clinical_metadata[
clinical_metadata["#Patient Identifier"].isin(count_data.index)
].copy()
clinical_metadata_subset.rename(
columns={
"#Patient Identifier": "ID_Sample",
"UICC Tumor Stage": "tnm_staging",
"Sex": "sex",
"Diagnosis Age": "age_at_sample",
"Overall Survival (Months)": "time_since_t0_death",
"Overall Survival Status": "status_os",
},
inplace=True,
)
clinical_metadata_subset["status_os"].replace(
{"1:DECEASED": 1, "0:LIVING": 0}, inplace=True
)
# Add the staging (< III, LS, IV then ES)
clinical_metadata_subset["staging"] = clinical_metadata_subset["tnm_staging"].apply(
lambda x: "ES" if x == "IV" else "LS" if x != "nan" else "LS"
)
try:
clinical_metadata_subset.set_index("ID_Sample", inplace=True)
except Exception as e:
print(f"Error setting index: {e}")
print(
f"Number of samples per stage: {clinical_metadata_subset.staging.value_counts()}")
We see that we have 72 patients that are in LS (limited stage, <= TNM stage III) and 9 ES (extensive stage, TNM stage IV).
And the count_data has this shape (zoom out if unable to see the entire table):
1
count_data
| B2M | FTL | HLA-B | ANXA1 | HLA-DRA | PFN1 | CTSD | CD74 | IGLL5 | LYZ | ... | CYP4F22 | NDST4 | CLEC12B | IL17C | AXDND1 | SCN2A | CNTNAP5 | RNF133 | DNAH8 | YAP1 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| sclc_ucologne_2015_S00022 | 2162.154403 | 2298.411925 | 1001.751408 | 23.633396 | 86.991391 | 678.728366 | 472.136116 | 27.912742 | 42.978858 | 28.775967 | ... | 2.750689 | 8.490031 | 2.725959 | 2.992372 | 2.715768 | 3.514570 | 8.136591 | 2.696780 | 2.728837 | 3.277595 |
| sclc_ucologne_2015_S00035 | 3160.088999 | 3745.273955 | 648.107862 | 102.795278 | 276.450852 | 615.955668 | 524.715629 | 50.000057 | 46.280251 | 84.170639 | ... | 2.192853 | 8.469487 | 2.246005 | 3.080709 | 2.148493 | 7.893646 | 2.475983 | 2.318928 | 2.155952 | 3.605534 |
| sclc_ucologne_2015_S00050 | 2052.014302 | 2118.938263 | 879.408577 | 36.147493 | 423.594166 | 532.336017 | 489.297038 | 109.557077 | 244.091360 | 212.364860 | ... | 3.146193 | 6.516990 | 2.984626 | 3.215373 | 3.042088 | 4.725947 | 18.991687 | 3.011319 | 3.134531 | 3.915876 |
| sclc_ucologne_2015_S00213 | 9640.167384 | 14605.657288 | 2289.823486 | 121.938662 | 2079.767782 | 884.827811 | 1074.222337 | 176.478753 | 5425.059394 | 2610.552043 | ... | 2.681442 | 15.147577 | 2.421056 | 3.124963 | 2.386332 | 4.895384 | 2.558535 | 2.451218 | 2.389190 | 4.352976 |
| sclc_ucologne_2015_S00356 | 5004.217605 | 4981.908336 | 835.222854 | 119.778900 | 983.900900 | 502.215145 | 514.830956 | 122.104755 | 1418.010653 | 499.754643 | ... | 2.829266 | 3.128775 | 2.719074 | 3.650863 | 2.776094 | 10.940923 | 4.448620 | 3.396203 | 2.740640 | 5.575360 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| sclc_ucologne_2015_S02375 | 6812.104434 | 11667.759405 | 1509.701805 | 1282.925898 | 2003.241765 | 1048.792635 | 1064.024598 | 183.230711 | 318.635008 | 572.531677 | ... | 2.093379 | 2.280902 | 2.106701 | 2.041678 | 2.030944 | 2.036819 | 3.085045 | 1.997111 | 2.072963 | 2.719448 |
| sclc_ucologne_2015_S02376 | 8251.028221 | 3238.868715 | 3740.925445 | 83.274742 | 501.582874 | 1147.828117 | 389.587217 | 58.089660 | 541.181302 | 293.667019 | ... | 2.179461 | 6.646879 | 2.105514 | 2.843782 | 2.154875 | 5.445269 | 6.453248 | 2.294543 | 2.117667 | 3.164042 |
| sclc_ucologne_2015_S02378 | 9520.367291 | 6724.043881 | 2958.637473 | 209.366759 | 1717.925348 | 677.814857 | 1175.556034 | 183.766750 | 267.717238 | 523.855194 | ... | 2.772599 | 2.877168 | 2.858701 | 3.147568 | 2.824992 | 2.806729 | 3.585936 | 2.965653 | 2.768039 | 6.153518 |
| sclc_ucologne_2015_S02382 | 3779.392094 | 2525.732381 | 577.241843 | 161.040659 | 226.311589 | 307.053940 | 236.202220 | 49.486211 | 557.569124 | 478.677608 | ... | 3.289202 | 3.953085 | 3.289202 | 6.149953 | 3.346079 | 4.930345 | 8.773256 | 3.703050 | 3.311947 | 7.840243 |
| sclc_ucologne_2015_S02397 | 4621.667938 | 5300.872063 | 1401.574063 | 82.634577 | 444.025069 | 779.949035 | 413.593408 | 72.070549 | 338.586292 | 227.066808 | ... | 2.991421 | 14.824241 | 3.086601 | 4.139198 | 2.982478 | 13.696145 | 4.118115 | 3.192017 | 2.981595 | 4.707087 |
81 rows × 1294 columns
Let’s now create an AnnData object and add the log_tpm_counts to it. This will be useful later on for plotting of the main TFs (transcription factors) for each group or subtype.
1
2
3
4
5
6
7
8
9
10
11
adata = ad.AnnData(count_data)
adata.obs = clinical_metadata_subset.copy()
adata.var = pd.DataFrame(count_data.columns)
adata.layers["tpm_counts"] = adata.X.copy()
adata.layers["log_tpm_counts"] = np.log1p(count_data.values).copy()
adata.var.set_index(0, inplace=True)
While one could run cNMF using all the genes from the bulk dataset, only a few thousand genes are needed to identify the subtypes. There are several methods to select the most relevant genes, but for this tutorial, we will use the discovery set from the original publication (Gay et al. 2021). These can be found in the supplementary material of the paper (see DOI above).
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
# Load genes 1300 discovery set
import openpyxl
final_genes = pd.read_excel(
_base_dir / "data" / "files" / "1-s2.0-S1535610820306620-mmc2.xlsx",
header=1,
)
final_genes = final_genes["Unnamed: 0"].tolist()
final_genes.append("YAP1") # adding YAP1 to this list just to test the expression of YAP1 across the samples to see that it shows a somewhat even expression across identified subtypes.
missing_genes = [gene for gene in final_genes if gene not in adata.var_names]
print("Missing genes:", missing_genes)
final_genes_filtered = [
gene for gene in final_genes if gene in adata.var_names]
final_adata = adata[:, final_genes_filtered].copy()
final_adata.X = final_adata.layers["tpm_counts"].copy()
Lets now crate the TPM counts file that will be used by cNMF.
1
2
3
4
5
6
counts_file_new = "./temp_counts_subtype_identification.txt"
pd.DataFrame(
adata.X,
index=adata.obs_names,
columns=adata.var_names,
).to_csv(counts_file_new, sep="\t")
Let’s now run cNMF. Here, we will test a range of components (from 2 to 8). This is generally done if you don’t yet know the optimal number of components/groups/subtypes to use. However, from the literature, we already know for SCLC that the field recognizes this number to be 4, so we will already use this number here.
Running cNMF
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
subtype_col_name="nmf-group-de-novo"
k: int = 4
density_threshold: float = 0.1
components: range = range(2, 8)
mandatory_genes: list = ["ASCL1", "NEUROD1", "POU2F3", "YAP1"]
n_iter: int = 150
total_workers: int = 1
num_highvar_genes: int = 2000 # number
ntop_genes: int = 5000
data_save_path: str = "./"
layer = "tpm_counts"
cnmf_obj_new = cNMF(
output_dir="./cnmf_results_selected_genes", name="nmf_selected_genes"
)
cnmf_obj_new.prepare(
counts_fn=counts_file_new,
tpm_fn=f"{_base_dir}/temp_counts_subtype_identification.txt", # note that we can load the TPM counts file directly. Generally, for this package, one can run the raw counts and then it will convert to TPM, but since we don't have access to the raw counts here, we will load the TPM counts directly, which have been calculated from the original FPKM values.
components=components,
n_iter=n_iter,
num_highvar_genes=num_highvar_genes, # The package may already use this number to use just the top 2000 genes that are highly variable across the samples. 2000 is the default number, but you can change it to something else. In this case, the total number of genes is around 1300, so this number being 2000 won't do much harm.
)
cnmf_obj_new.factorize(worker_i=0, total_workers=total_workers)
cnmf_obj_new.combine()
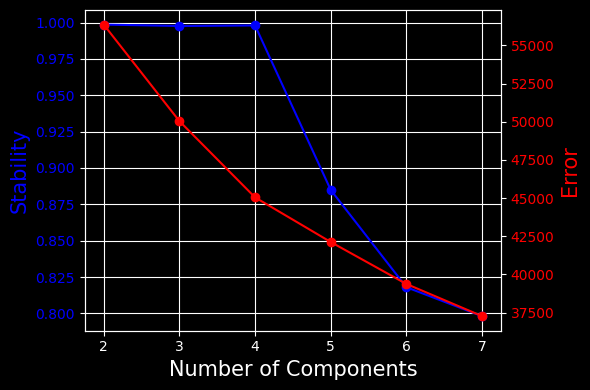
This plot will show the stability and error for each of the K tested (in our case, from 2 to 7 components). We see that the optimal number of components is 4, which offers the best stability and lowest error, which once again confirms the literature.
1
2
3
4
5
6
7
8
cnmf_obj_new.consensus(
k=k,
density_threshold=density_threshold,
show_clustering=True,
close_clustergram_fig=False,
)
cnmf_obj_new.k_selection_plot()
1
2
3
usage_matrix_nmf_var, gep_scores_nmf_var, gep_tpm_nmf_var, topgenes_nmf_var = (
cnmf_obj_new.load_results(K=k, density_threshold=density_threshold)
) # capture the usage matrix, GEP scores, etc.
Now that we have calculated the usage matrix, we can merge it with the original AnnData object. This is a normalized value, that sums to 1 across our selected K. In this case, we will have 4 new columns, from 1 to 4, representing each of the 4 components.
We can also then take the maximum value of the usage matrix, and give a categorical assignment to each sample based on the maximum value. This is useful for other downstream analysis, such as differential expression analysis, boxplot visualizations, etc.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
adata_nmf = adata.copy()
adata_nmf.obs = pd.merge(
left=adata_nmf.obs,
right=usage_matrix_nmf_var,
how="left",
left_index=True,
right_index=True,
)
top_genes_nmf = []
print(f"Selecting top {ntop_genes} genes for each subtype")
for gep in gep_scores_nmf_var.columns:
top_genes_nmf.append(
list(
gep_scores_nmf_var.sort_values(by=gep, ascending=False).index[
:ntop_genes
]
)
)
top_genes_nmf = pd.DataFrame(top_genes_nmf, index=gep_scores_nmf_var.columns).T
for col in top_genes_nmf.columns:
print([g for g in top_genes_nmf[col] if g in mandatory_genes])
adata_nmf.obs[subtype_col_name] = usage_matrix_nmf_var.idxmax(
axis=1
).astype(str)
print(
f"Count of groups: {adata_nmf.obs[subtype_col_name].value_counts()}"
)
gene_df = pd.DataFrame({"gene": adata_nmf.var_names.tolist()})
print(
f"Saving top {ntop_genes} NMF genes to {data_save_path + 'nmf_genes.csv'}"
)
gene_df.to_csv(data_save_path + "nmf_genes.csv")
adata_nmf.obs[["nmf-group-de-novo",1,2,3,4]]
Our adata.obs now has the subtype scores and the nmf-group-de-novo which is essentially the maximum of the subtype scores, so we can associate a subtype for each sample (zoom out if unable to see the entire table). Doing a simple .value_counts() on this column will give us the number of samples per subtype which goes as follows:
| Index | Value |
|---|---|
| 1 | 29 |
| 2 | 21 |
| 3 | 18 |
| 4 | 13 |
Additionally, the subtype scores, as these are normalized, they can be interpreted as proportions of a given subtype across each sample. This is particularly useful, as we can do correlations with other biomarkers we may have, or to simply indicate the proportion of subtypes for each sample since, realistically, one patient is likely to have multiple subtypes at once, within a single sample.
One could also combine these scores and define thresholds to define “mixed” subtypes (e.g. A/N, A/I, etc). This can be particularly useful if we want to be a bit more specific about the subtypes we define, especially in SCLC, as research suggests that the subtypes are not always clear-cut
| nmf-group-de-novo | 1 | 2 | 3 | 4 | |
|---|---|---|---|---|---|
| ID_Sample | |||||
| sclc_ucologne_2015_S00022 | 2 | 0.472633 | 0.506275 | 0.006064 | 0.015028 |
| sclc_ucologne_2015_S00035 | 1 | 0.836964 | 0.000000 | 0.106799 | 0.056237 |
| sclc_ucologne_2015_S00050 | 2 | 0.174711 | 0.798140 | 0.027149 | 0.000000 |
| sclc_ucologne_2015_S00213 | 3 | 0.417614 | 0.000000 | 0.539206 | 0.043180 |
| sclc_ucologne_2015_S00356 | 1 | 0.427478 | 0.282972 | 0.198701 | 0.090849 |
| ... | ... | ... | ... | ... | ... |
| sclc_ucologne_2015_S02375 | 4 | 0.034423 | 0.000000 | 0.384087 | 0.581490 |
| sclc_ucologne_2015_S02376 | 1 | 0.656548 | 0.029480 | 0.260617 | 0.053356 |
| sclc_ucologne_2015_S02378 | 3 | 0.175559 | 0.215421 | 0.337799 | 0.271221 |
| sclc_ucologne_2015_S02382 | 2 | 0.228773 | 0.581242 | 0.023989 | 0.165996 |
| sclc_ucologne_2015_S02397 | 1 | 0.622625 | 0.079786 | 0.144879 | 0.152710 |
81 rows × 5 columns
Plotting the results
Now that we have our subtypes, it is time to plot some of the known markers from SCLC and see which group has highest expression for each of the key markets. To explore this, let’s do some boxplots, showing the groups (i.e. subtypes) against key genes using the log_tpm_counts that we calculated earlier.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
# Start by creating a dataframe with the log_tpm_counts and the ID_Samples
key_markers_boxplot = pd.DataFrame(
adata_nmf.layers["log_tpm_counts"],
columns=adata_nmf.var_names,
index=adata_nmf.obs_names,
)
# Subset just the key genes
key_markers_boxplot = key_markers_boxplot[["ASCL1", "NEUROD1", "POU2F3", "YAP1"]]
# Add the NMF grouping we calculated earlier using cNMF
merged_data = key_markers_boxplot.merge(adata_nmf.obs[["nmf-group-de-novo"]], on="ID_Sample", how="inner")
# Now, plot each of the key genes against the NMF groups
for key_genes in ["ASCL1", "NEUROD1", "YAP1", "POU2F3"]:
ax = sns.boxplot(x="nmf-group-de-novo", y=key_genes, data=merged_data, palette="Set2")
ax.set(xlabel="NMF groups", ylabel="log TPM counts")
ax.set_title(f"Boxplot of {key_genes} expression per NMF group", fontsize=20)
ax.set_ylabel("log TPM counts")
ax.set_xlabel("NMF groups")
plt.show()
plt.close()
This will give the following boxplots:
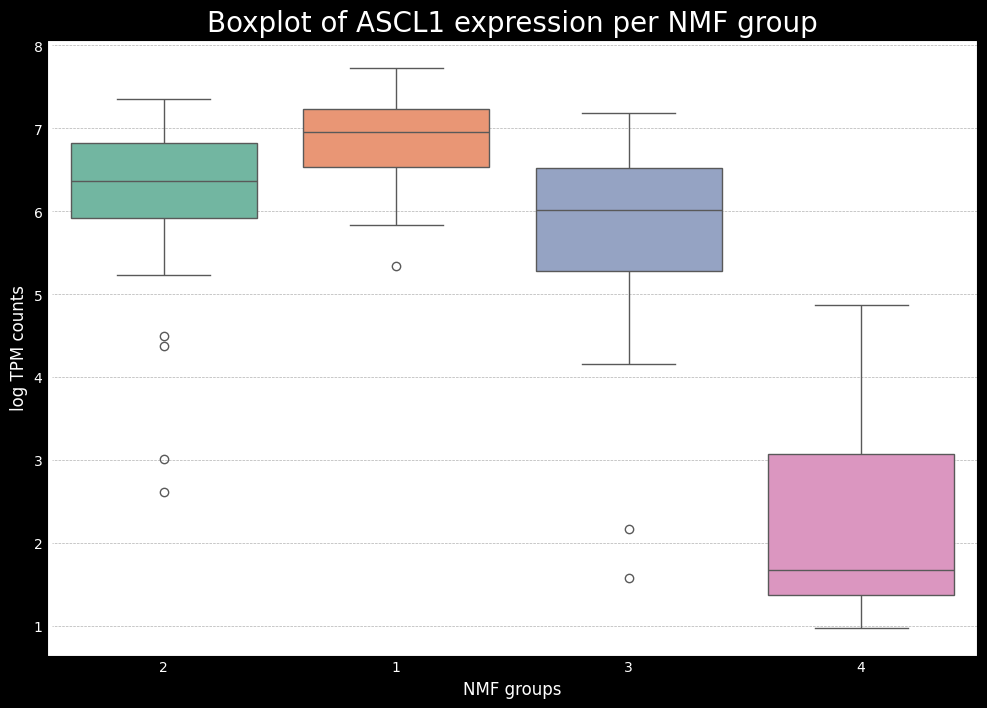
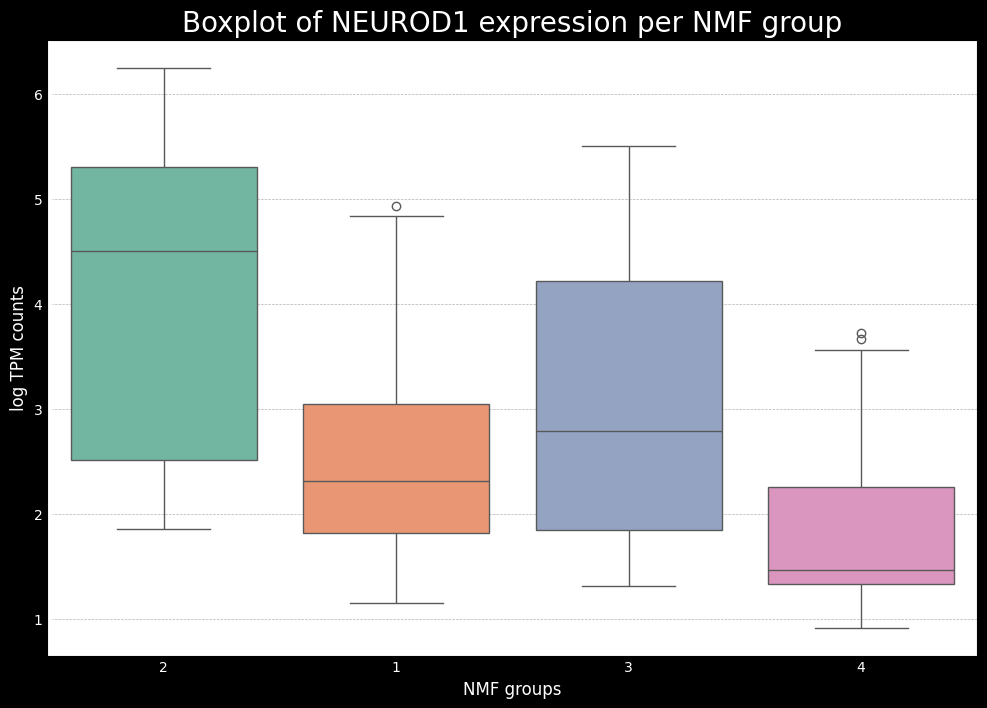
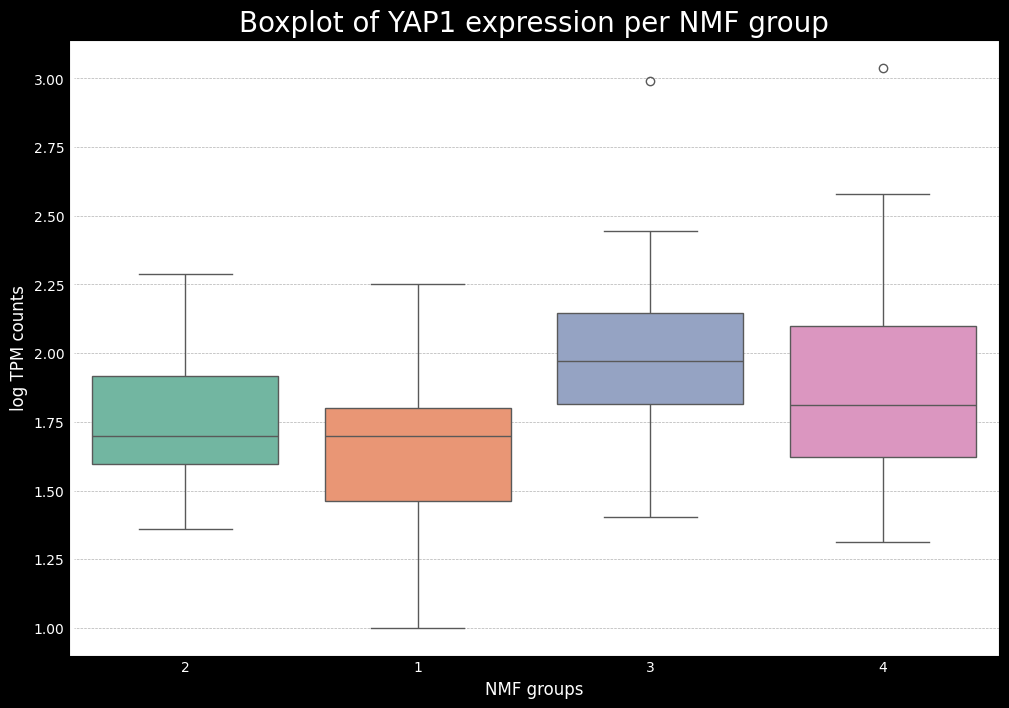
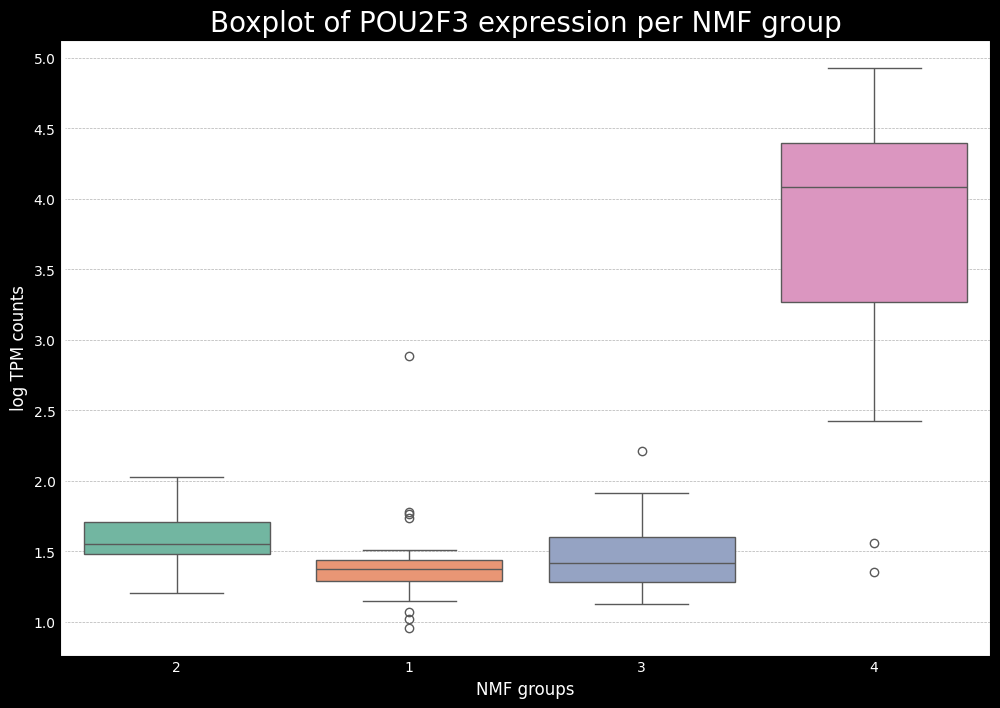
This gives us a good idea of which subtypes are associated with each of the key genes. Now, ideally, if we have the raw counts, the next step is to do some differential expression analysis to see which of the key markers from SCLC (or another cancer being analyzed) are differentially expressed between the subtypes. To do this, one can use the pydeseq2 package, which is a popular tool for differential expression analysis in python. While it is not the focus of this tutorial, I will bring another tutorial later on showing an example of how to use pydeseq2 and a modification to do both one-vs-one and one-vs-all comparisons.
The next step is to do some more visualizations that will help us understand the subtypes better, this time using gene sets and a radial plot. The gene sets we will use (see code below) are NE (neuroendocrine), nonNE (non-neuroendocrine), T-eff (tumor-effector), B/PC (B-cell and plasma cell), and APM (antigen processing and presentation) and Checkpt (immune checkpoint).
Additionally, we will also use a mapping between the numerical NMF groups, and the subtypes based on the boxplots made earlier. We see that cluster 1 has higher expression for ASCL1, so we associate this cluster to A subtype. Cluster 2 has higher expression for NEUROD1, so we associate this cluster to N subtype. Cluster 4 has higher expression for POU2F3, so we associate this cluster to P subtype. The last cluster, cluster 3 can, can be associated with the Inflammed subtype. We see that the expression of YAP1 is not sufficient to define a subtype on its own.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
adata_analysis_radial = adata_nmf.copy()
adata_analysis_radial.obs = adata_analysis_radial.obs[
[
1,
2,
3,
4,
"nmf-group-de-novo",
]
]
# Since we already have the log_tpm_counts, we can use them directly, and then only scale them
adata_analysis_radial.X = adata_analysis_radial.layers["log_tpm_counts"].copy()
sc.pp.scale(adata_analysis_radial)
# These will be the gene sets we will use
gene_categories = {
"NE": ["CHGA", "DLL3", "NEUROD1", "INSM1", "ASCL1"],
"nonNE": ["YAP1", "POU2F3", "MYC", "REST"],
"T-eff": [
"CD8A",
"GZMA",
"GZMB",
"PRF1",
"IFNG",
"CXCL9",
"CXCL10",
"TBX21",
],
"B/PC": ["CD79A", "MS4A1", "MZB1", "JCHAIN"],
"APM": ["TAP1", "TAP2", "B2M", "HLA-A", "HLA-C"],
"Checkpt": ["PDCD1", "CD274", "LAG3", "CTLA4", "BTLA", "TIGIT"],
}
# Compute scores for each category
for category, genes in gene_categories.items():
sc.tl.score_genes(adata_analysis_radial, genes,
score_name=f"{category}_score")
# Calculate mean scores for each NMF group
nmf_scores = adata_analysis_radial.obs.groupby("nmf-group-de-novo").mean()
# Normalize scores (Z-score across subsets)
score_columns = [f"{cat}_score" for cat in gene_categories.keys()]
nmf_scores_norm = (
nmf_scores[score_columns] - nmf_scores[score_columns].mean()
) / nmf_scores[score_columns].std()
# Define NMF labels
nmf_labels = {
"1": "A",
"2": "N",
"3": "I",
"4": "P",
}
# Colors for each NMF group
colors = ["#8A7EB5", "#67C28E", "#E75480", "#FFA500"]
color_map = {
"1": "#4169E1", # A - RoyalBlue (cold)
"2": "#008080", # N - Teal (cold)
"3": "#DC143C", # I-nNE - Crimson (warm)
"4": "#FF8C00", # I-NE - DarkOrange (warm)
}
fig = plt.figure(figsize=(8, 8))
ax = plt.subplot(111, projection="polar")
# Prepare angles
categories = list(gene_categories.keys())
N = len(categories)
angles = [n / float(N) * 2 * np.pi for n in range(N)]
angles += angles[:1] # Complete the loop
# Plot each group
for nmf_group in sorted(nmf_scores_norm.index):
data = nmf_scores_norm.loc[nmf_group, score_columns].to_frame().T
values = data.values.flatten().tolist()
values += values[:1] # Complete the loop
label = nmf_labels.get(nmf_group, nmf_group)
color = color_map[nmf_group]
ax.plot(
angles,
values,
linewidth=2.5,
linestyle="solid",
label=f"NMF{nmf_group}: {label}",
color=color,
)
ax.fill(angles, values, color=color, alpha=0.2)
# Format the plot
ax.set_theta_offset(np.pi / 2)
ax.set_theta_direction(-1)
ax.set_xticks(angles[:-1])
ax.set_xticklabels(categories, size=15)
ax.set_yticklabels([]) # Hide radial labels
ax.grid(True, color="grey", alpha=0.3)
ax.spines["polar"].set_visible(False)
ax.legend(loc="upper right", bbox_to_anchor=(1.3, 1.1), fontsize=12)
ax.set_title("Radial plot of NMF groups", size=20, y=1.1)
plt.tight_layout()
plt.show()
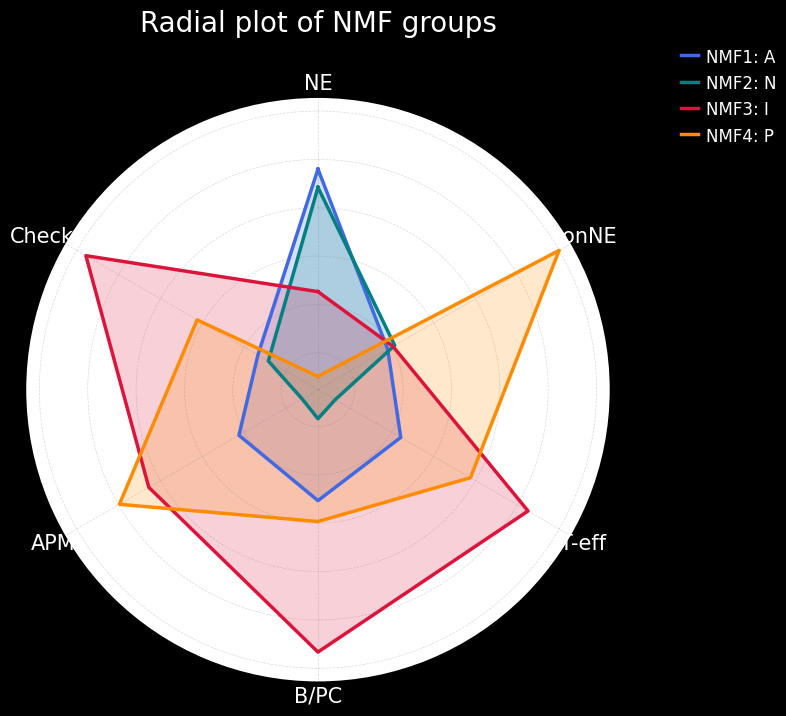
Conclusion
And there you have it! We have now identified the molecular subtypes of SCLC using cNMF, with matching distributions and TFs expression profiles, compared to the original publication.
There is a lot we can do with this information. Cell type deconvolution on the bulk RNA data to further evaluate cell type composition, survival analysis, differential expression analysis, functional enrichment analysis, etc. I plan on bringing more tutorials across all things related to transcriptomics and machine learning, so stay tuned!
Enjoy Reading This Article?
Here are some more articles you might like to read next: